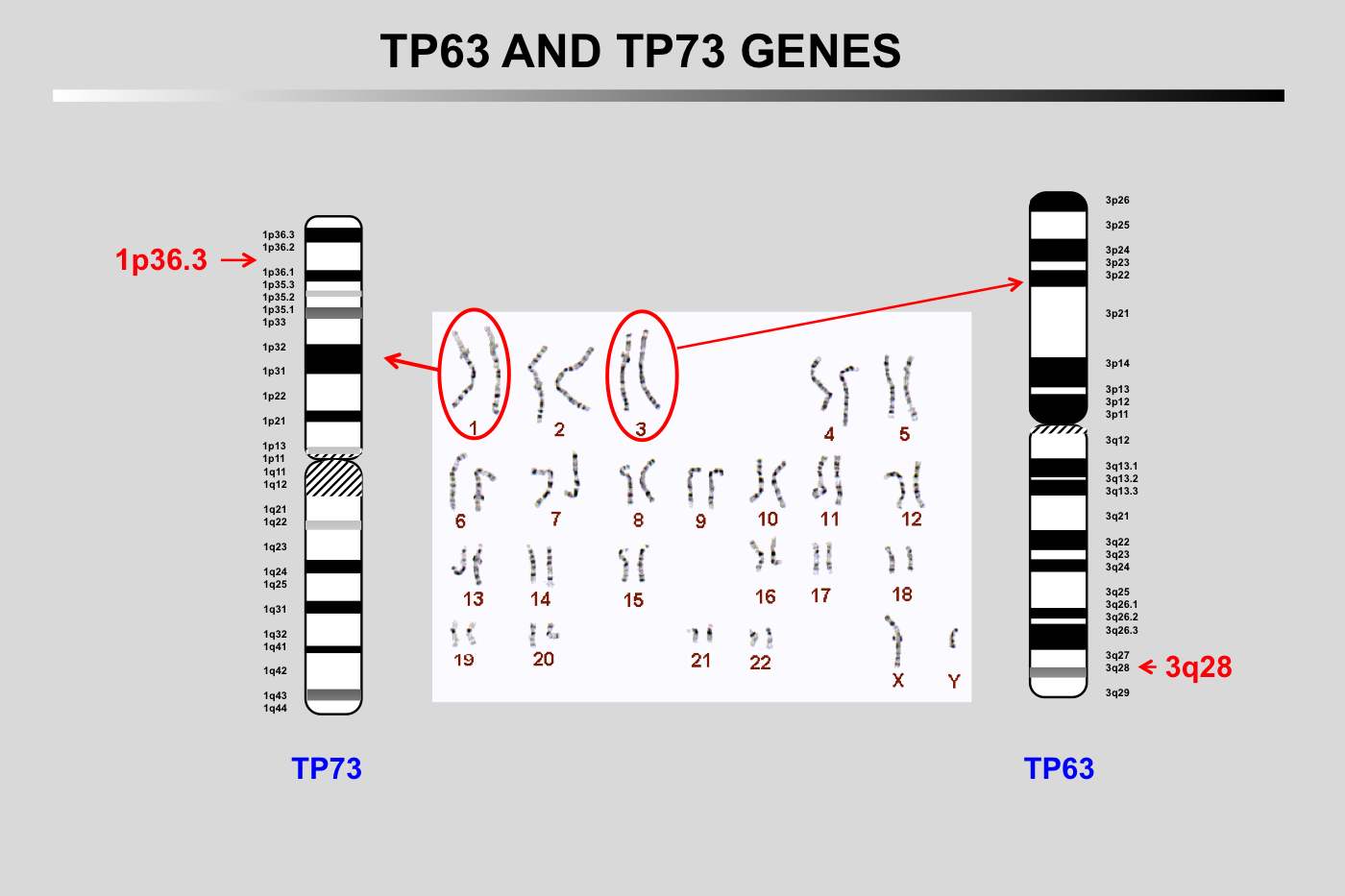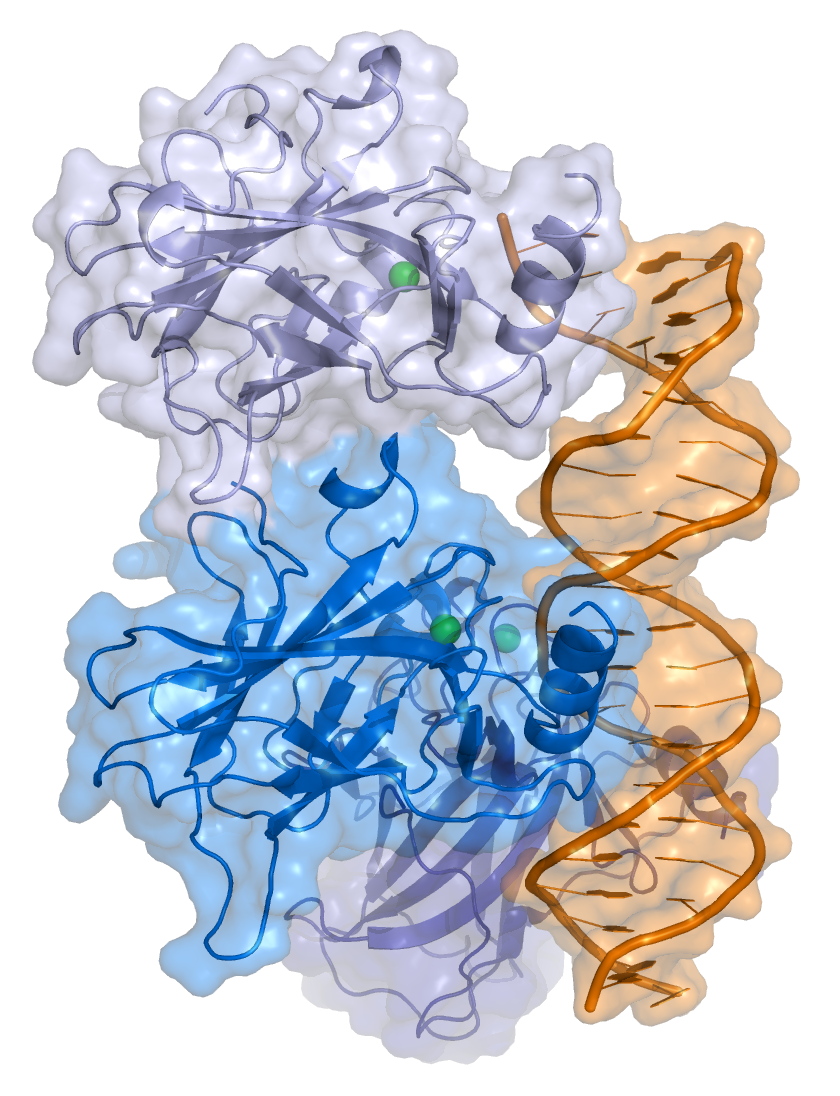
TP53 is a sequence-specific transcription factor that binds to specific response elements, which comprise two half sites of the nucleotide sequence RRRCWWGYYY (in which R = purine, W = A or T, and Y = pyrimidine), typically separated by a 0–13 nucleotide spacer.
The major TP53 isoforms (TP53 alpha or P1, 393 aa) comprise several domains that are responsible for sequence-specific DNA binding, transcriptional activation and tetramerization. TP53 activity is regulated by multiple post-translational modifications.
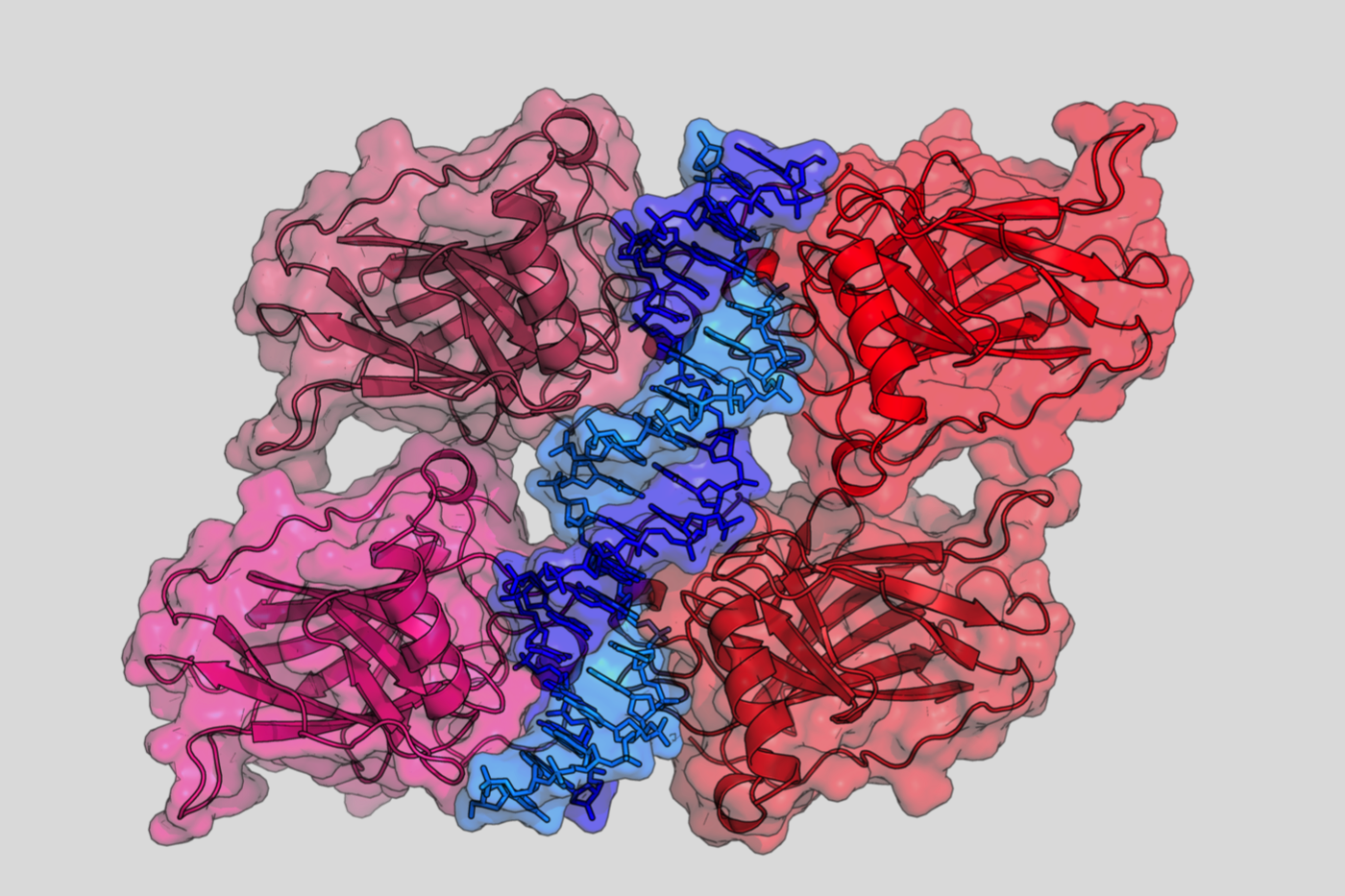
Crystal structure of four p53 DNA binding domains
(http://en.wikipedia.org/wiki/P53)
Click on the links below for further details
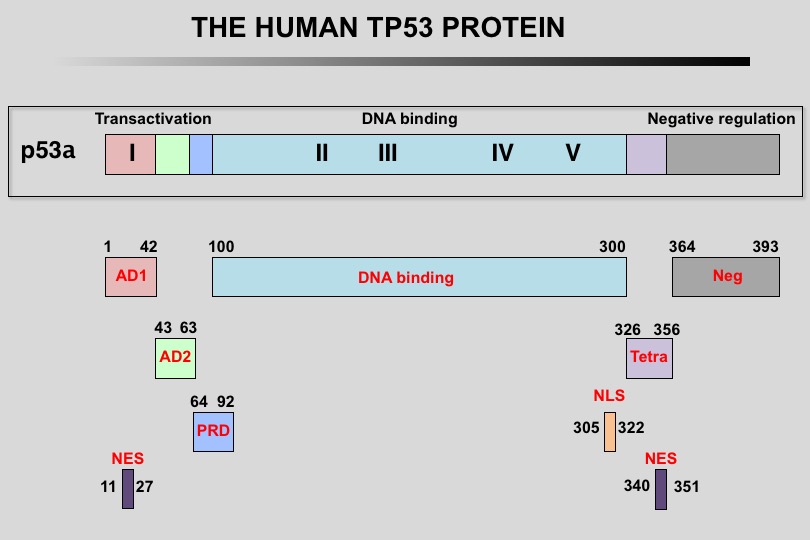
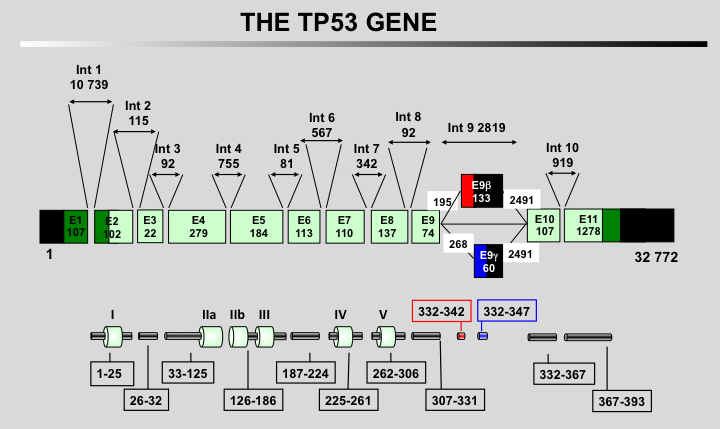
The human TP53 protein: Human TP53 protein can be divided into multiple regions, each corresponding to specific functions:
1) The amino-terminus contains two independent transactivation domains (aa 1-42 and 43 to 63) and the mdm2 protein binding site (aa 13-29). It also contains the Highly Conserved Domain I (HCD I). A nuclear exclusion signal (NES) has also be localized in the amino-terminus of the protein.
2) Region 64-92 (PRD, Proline-Rich Domain) contains a series of repeated proline residues that are conserved in the majority of TP53 proteins.
3) The central region (101-300) contains the DNA binding domain. It is the target of 85% of TP53 mutations found in human cancers. It also includes HCD II to V.
4) The oligomerization domain (307-355, TET) consists of a beta-strand, followed by an alpha-helix necessary for dimerization, as TP53 is composed of a dimer of two dimers. A nuclear export signal (NES) is localized in this oligomerization domain.
5) The carboxy-terminus of p53 (356-393) contains 3 nuclear localization signals (NLS) and a nonspecific DNA-binding domain. This region is also involved in downregulation of DNA binding of the central domain.
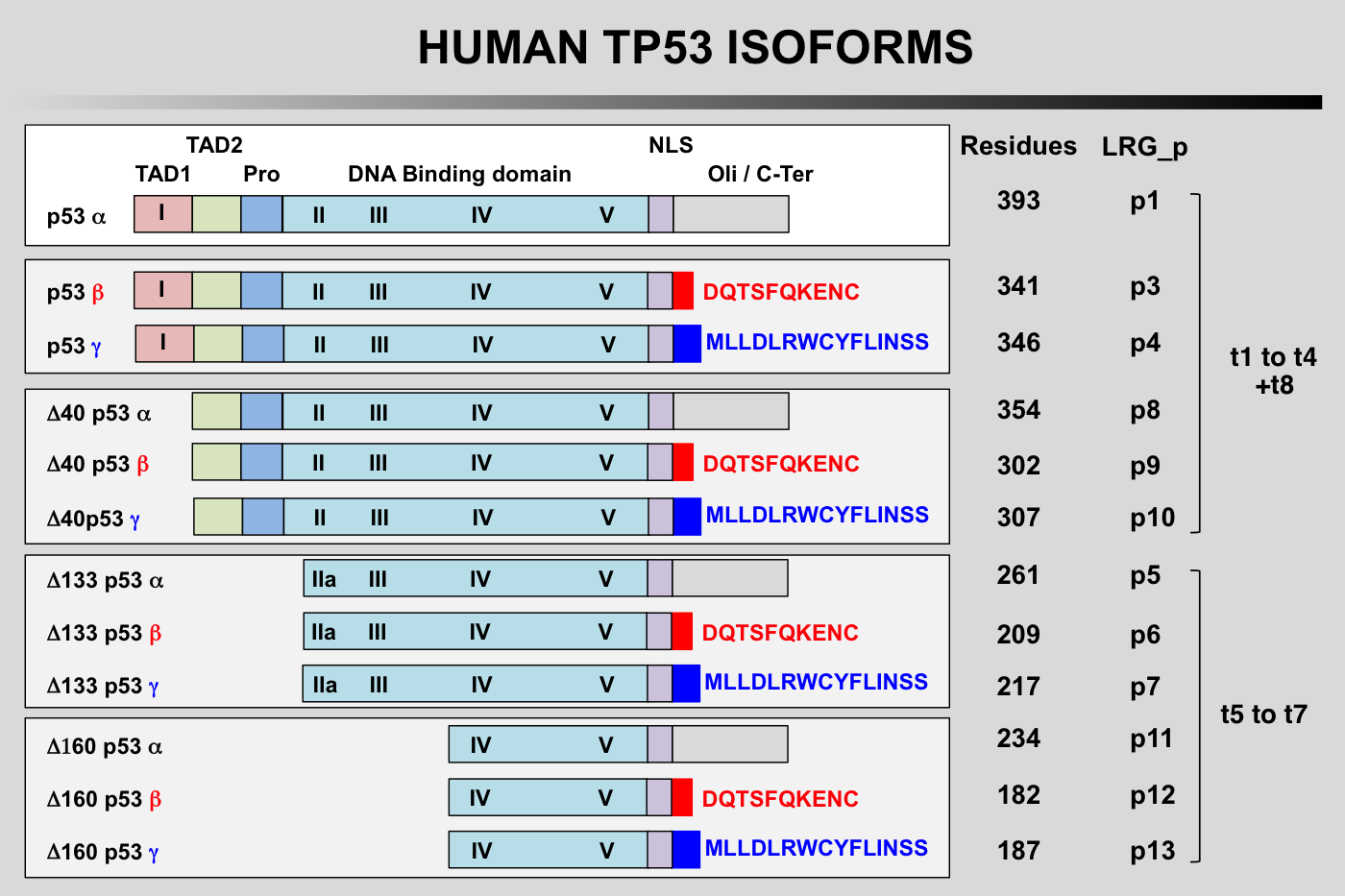
TP53 isoforms nomenclature according to the LRG
TP53 Isoforms:
The first group of TP53 is translated from the five P1-initiated TP53 mRNAs t1-t4 and t8. They can begin at codon 1 or codon 40 leading to the expression of either full-length TP53 protein (TP53) or truncated TP53 protein missing the first transactivation domain (Delta40p53). Furthermore, Delta40p53 proteins can also be encoded by alternatively-spliced TP53 mRNA t8 still containing intron 2. It is important to note that such a transcript encodes only Delta40p53 because intron 2 contains a stop codon that prevents expression of full-length p53.
The second group with different amino-termini contains TP53 isoforms encoded by three novel mRNAs, t5, t6 and t7. These transcripts are produced by the P2 promoter localized in intron 4 and probably extending to exon 3 and start from a novel transcription initiation site at the 3' end of intron 4. They generate TP53 proteins with different amino-termini starting at either amino acid 133 or 160 (Delta133p53 and Delta160p53).
Finally, each of the different amino-termini can be combined with a different carboxy-terminus.
The alternatively spliced transcripts containing exon 9 beta (t3 and t6) each use two translation start sites to encode 4 TP53 isoforms combining different amino-termini with the beta carboxy-terminus (TP53β Delta40p53β Delta133p53βand Delta160p53β). Similarly, t4 and t7 containing exon 9 gamma encode 4 TP53 isoforms combining different amino-termini with the gamma carboxy-terminus (TP53γ Delta40p53γ Delta133p53γand Delta160p53γ. Although physiological expression of intron 2 containing t8 equivalents with exons 9 beta and 9 gamma has yet to be demonstrated experimentally, these transcripts would probably encode the existing Delta40p53β and Delta40p53γisoforms.
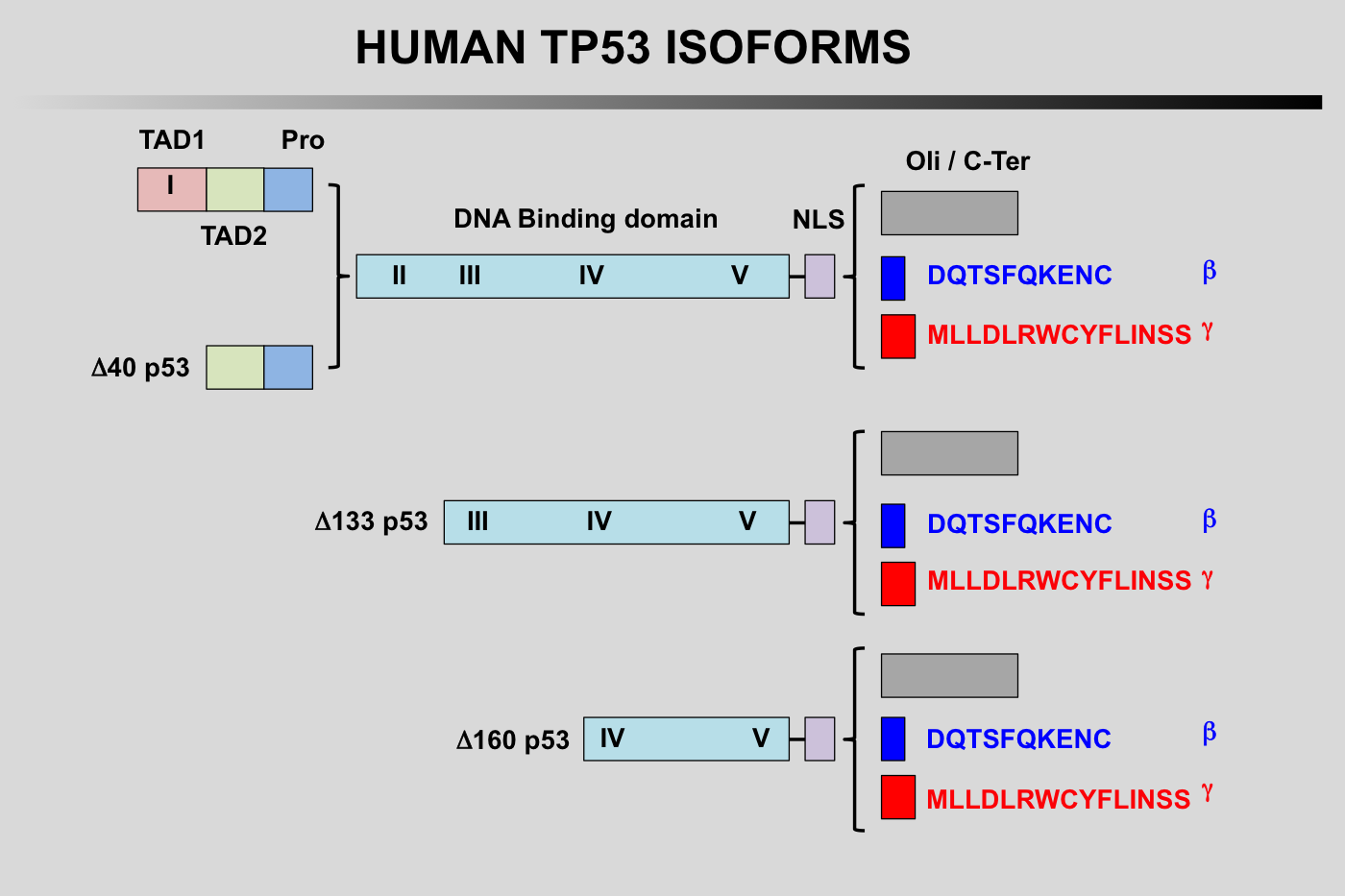
TP53 isoforms
A novel TP53 isoform, TP53 psi, have recently been identified. This protein is generated by the alternative 3' splice site in intron 6 leading to the synthesis of a truncated protein at residue 224 which is unable to bind to DNA (Senturk et al. 2014).
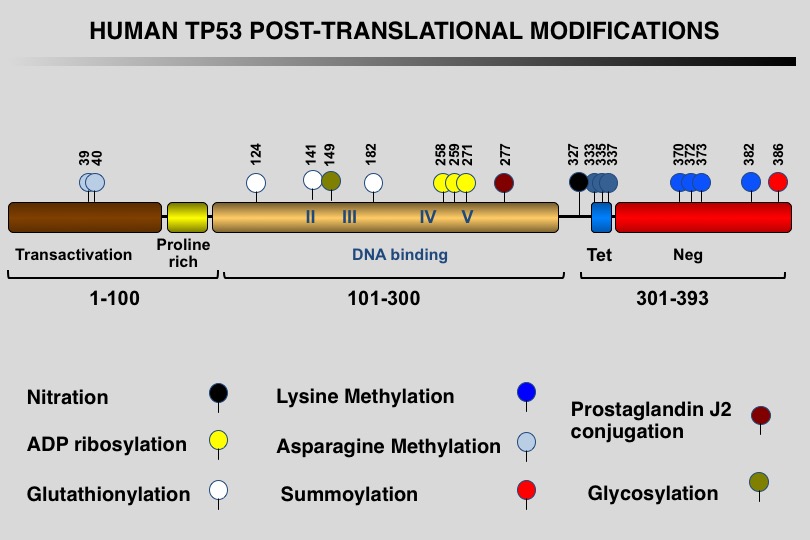
Like many eukaryotic transcription factors, the TP53 protein is the target of multiple post-translational modifications (PTM) that modulate its activity (Meek and Anderson, 2009; Nguyen et al. 2014).
Twelve different types of modification targeting 62 of the 393 residues of the TP53 protein have been identified, the most frequent being phosphorylation and ubiquitination.
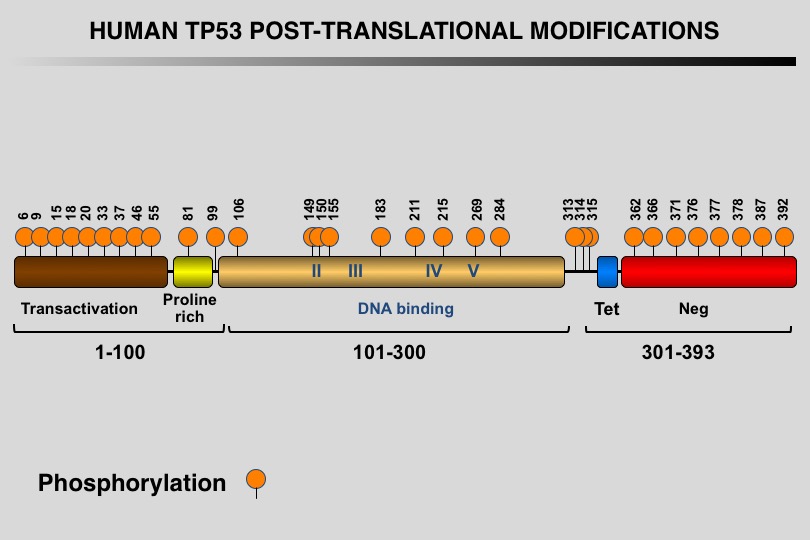
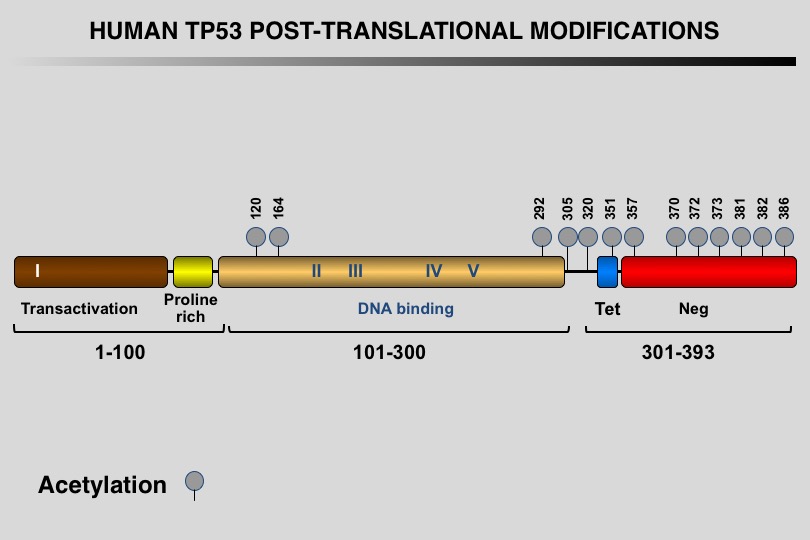
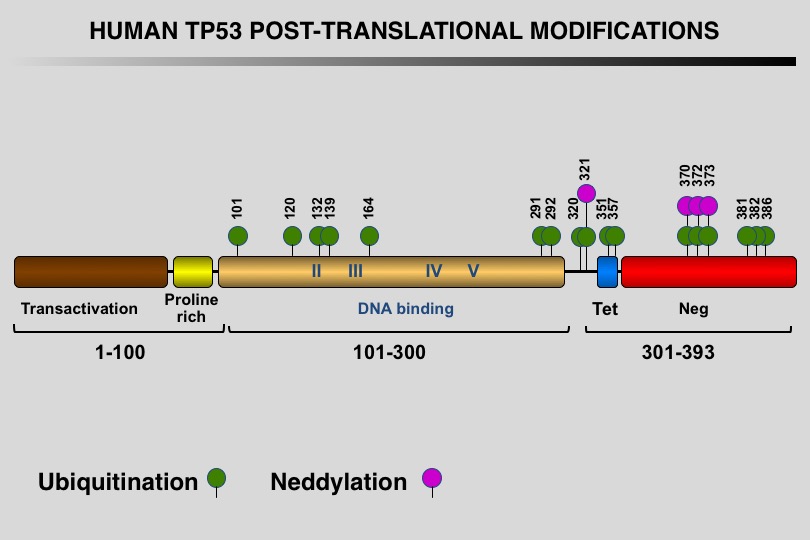
Meek DW, Anderson CW (2009) Posttranslational modification of p53: cooperative integrators of function. Cold Spring Harb Perspect Biol 1: a000950. 20457558
Nguyen TA, Menendez D, Resnick MA, Anderson CW (2014) Mutant TP53 Posttranslational Modifications: Challenges and Opportunities. Hum Mutat 35: 738-755. 24395704
Hock AK, Vousden KH (2014) The role of ubiquitin modification in the regulation of p53. Biochim Biophys Acta 1843: 137-149. 23742843
Kruse JP, Gu W (2008) SnapShot: p53 posttranslational modifications. Cell 133: 930-30.e1. 18510935
Khoury MP, Bourdon JC (2011) p53 Isoforms: An Intracellular Microprocessor? Genes Cancer 2: 453-465. 21779513
DeHart, C. J., Chahal, J. S., Flint, S. J., and Perlman, D. H. (2014). Extensive post-translational modification of active and inactivated forms of endogenous p53. Mol Cell Proteomics 13, 1-17.