The full FAQ can be downoad here
Last modified: October 2017
i) Mutations published in peer-reviewed journals. Only unambiguous mutations described with accurate coordinates are included. Imprecise mutations such as "exon 6, C>T" or "splice in intron 5" have been omitted. Similarly, frameshift mutations such as "codon 216del" or R213fs" have also been omitted, as the size of the insertion and/or deletion is not included
Publications with a high error rates (incorrect correspondence between codon and amino acids or erroneous coordinates) have also been omitted.
ii) Published and unpublished data from two sequencing projects (ICGC and TCGA) have also been included.
https://icgc.org/
http://www.cbioportal.org/index.do
iii) For cell line data, in addition to publications, we have also included data from the cosmic database (http://cancer.sanger.ac.uk/cell_lines, the Cancer Cell Line Encyclopedia (CCLE, http://www.broadinstitute.org/ccle/home) and the NCI (http://discover.nci.nih.gov/cellminer/)
The current version of the UMD database uses the stable NCBI sequence NG_017013.2 as a reference for TP53 as well as hg18 (NCBI Build 36.1), hg19 (GRCH37) or hg38 (GRCh38) genome builds. This is a key issue as it will alleviate any problems associated with the use of multiple genome references by the various NGS pipelines.
The functional organization of the TP53 gene is more complex than previously thought: the NCBI’s RefSeq database now contains 15 different pairs of TP53 transcript and protein records references due to policy to associate only one RNA species to a single protein. Thus, several mRNA species encoding more than one protein have been duplicated with different RefSeq NM-accession numbers and two protein isoforms are represented by multiple RefSeq NP-accession numbers. To solve this confusing situation, TP53 specialists have joined forces with the Locus Reference Genomic (LRG) Consortium, which provides stable reference sequences and a coordinate system for permanent and unambiguous reporting of disease-causing variants in genes related to any pathology
The joint effort resulted in a recently released stable TP53 reference sequence, LRG_321 containing the genomic sequence from human genome build GRCh37.p13 (ftp://ftp.ebi.ac.uk/pub/databases/lrgex/LRG_321.xml). We believe that its annotation with precise labels and coordinates of 8 different TP53 transcripts (t1 to t8) and 12 isoforms (p1 and p3 to p13) will be preferred to the RefSeq identifier pairs provided by the NCBI for genome build GRCh37.p13.
Therefore, we have used LRG transcript and protein isoform numbers in the database.
Your publication could be missing for the following reasons:
• We may have failed to detect your publication: please send us a reprint and we will include your data.
• Your data may have already been reported in another publication. Please check the entire list of publications for other publications from your group. In the case of duplicate data, we usually include the older publication.
• The nomenclature used in the manuscript to describe the mutations may have been too ambiguous. If you like, send us a full description of the mutations using the official nomenclature and we will include your data.(http://www.hgvs.org/mutnomen/). You can use mutalyzer (https://mutalyzer.nl/) to check the accuracy of your mutations.
• There may have been too many erroneous mutations in your manuscript. Publications with more than 20% of errors (codon/amino acids/events) were discarded. If you like, send us a full description of the mutations using the official nomenclature and we will include your data.
Some of these mutations may already have been published. Please check the entire list of publications. If this is a mistake, let us know and we will update the database.
Check the description of these mutations, as they may have been too ambiguous to be included in the database.
The official nomenclature for the description of sequence variants is described by the Human Genome Variation Society (HGVS) (http://www.hgvs.org/mutnomen/). All mutations included in the UMD TP53 database have been carefully analysed and manually reviewed with mutalyzer (https://mutalyzer.nl/), a powerful tool developed by Jeroen F.J. Laros in close collaboration with HGVS.
We have noticed that numerous insertions and deletions were not accurately described (see some examples below). All these mutations have been carefully curated and modified to comply with the official nomenclature.
Example:
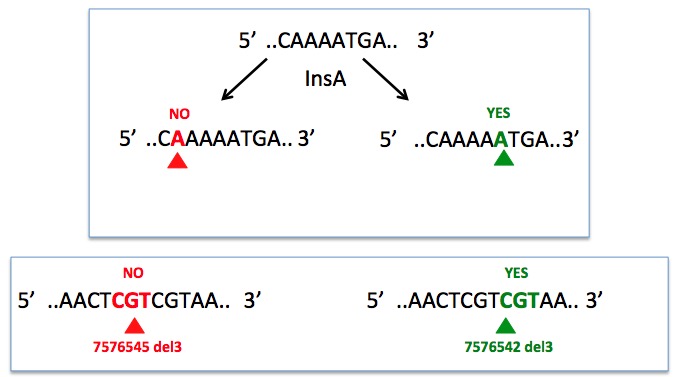
Many insertions described in the literature are in fact nucleotide duplications. This has also been correted in the database.
The UMD TP53 database only records the diversity of mutational events targeting the TP53 gene.
The variant MN_000546.5:c.1010G>A (NP_000537.3:p.R337H) is founder mutation that occurred in only one individual (or perhaps a few individuals) and has been transmitted to the entire population of descendents in Brazil. Therefore, only a single genetic event occurred in the population leading to dissemination of the mutation.
(see http://en.wikipedia.org/wiki/Founder_effect).
Defining this mutation as a hot spot mutation because it has been found in several hundred families is not correct.
Similarly, identical mutations found in matched primary tumours/metastases or preneoplastic lesions/primary tumours have been entered into the database only once, using the earliest lesion for disease category.
Similarly, identical germline mutations found in different individuals from the same family have been entered into the database only once,
Although the mutations have been very carefully entered into the database, a few errors may have been made. Please contact us in order to correct the database.
Send us a full description of the mutations using the official nomenclature and we will include your data.
Please contact us and we can discuss this issue, which could be of particular interest to improve curation of the database.
We have developed a curation procedure based on various types of statistical analysis of the database. Each publication has been assigned with an outlier index ranging from 0.1 to 15 with a cut-off of 2 (see for more info). Any value greater than 2 suggests that the publication should be considered to be suspicious and should be interpreted cautiously. 136 publications with an index ranging from 2 to have been identified and labelled as outliers. The few publications known to include a majority of sequencing artefacts have an index ranging between 5 and 12.
These publications have been kept in the database to provide the scientific community with access to an uncurated database with the possibility of developing and improving curation algorithms.
If you want to work with a curated database, you can easily filter out all publications labelled as outliers (see the read me file)
No problem. Contact us and we will make the requested changes. This email address is being protected from spambots. You need JavaScript enabled to view it.
Artefactual mutations from DNA extracted from paraffin-embedded tissue.
Duplicated series of TP53 mutations published in different journals.
Passenger TP53 mutations.
Unfrequent germline TP53 mutations.
Please use the latest publication and include the url of the website.
Leroy, B., Anderson, M., and Soussi, T. (2014). TP53 mutations in human cancer: database reassessment and prospects for the next decade. Hum Mutat 35, 672-688.
Pubmed access
Thank you.
No problem. Contact us so that we can provide you with the latest version of the database. Do not forget to mention the origin of the database in your publication. This email address is being protected from spambots. You need JavaScript enabled to view it.
Please contact us for further discussion. This email address is being protected from spambots. You need JavaScript enabled to view it.
Please contact us for further discussion. This email address is being protected from spambots. You need JavaScript enabled to view it.
This is an important issue, particularly if it is a germline variant.
If it is a frameshift variant (germline or somatic).
If the mutational event has been verified*, finding a novel deletion or insertion is not infrequent and the mutation could be considered to be pathogenic.
| Mutation type | Tissus | Interpretation |
| Germline variant* | Peripheral blood lymphocytes | Sequencing a larger number of individuals may allow identification of excessively rare, novel, non-pathogenic SNP. Until the pathogenicity of the variant has been clearly demonstrated (segregation with the disease or experimental analysis), this variant should be considered to be a Variant of Unknown Significance.
Various tools can be used to predict the pathogenicity of the mutation, but their sensitivity and specificity for TP53 are low. Use with caution. The effect on RNA translation or splicing is not included in predictive software. You can contact us for further discussion concerning this variant. |
| Somatic variants*, ** | Frozen tissues | In the absence of any functional information, there is no way to define whether this variant is a driver or a passenger mutation. |
| Somatic variants*, ** | Paraffin-embedded tissues | Sequencing of DNA from paraffin-embedded tissues is known to be associated with a high number of artefactual single-nucleotide changes (C:G>T:A). Careful control is necessary to validate this variant.
In the absence of any functional information, there is no way to define whether this variant is a driver or a passenger mutation. |
* Assuming that sequencing of the genetic material has been carefully performed and controlled.
** It is assumed that the somatic origin of this variant has been validated by sequencing normal DNA from the same individual.
